Rewiring oncogene enhancers and 3D genome organisation in primary cancers
New findings from team eDyNAmiC
Meet the global network of early-career researchers driving the development of cancer’s most intricate maps, as part of our 3D Tumour Mapping challenge.
The tumour ecosystem, a web of interactions between the tumour’s microenvironment and the rest of the body, evolves over time, producing a highly interconnected structure of cancer cells, normal tissues, immune cells and structural support. A detailed 3D map of this complex ecosystem – including each cell’s type, location and genetic makeup, and, importantly, how it interacts with and influences its neighbours – would reveal hidden information about tumour biology. This map could guide researchers in search of novel ways to diagnose and treat cancer, and prevent metastasis or recurrence.
The Cancer Grand Challenges IMAXT team, in taking on the 3D Tumour Mapping challenge, set out to develop a new way to create a map of the molecular and cellular tumour environment, through an integrated pipeline of techniques, which can be explored in virtual reality. The power of their approach lies in the integration of these techniques – each powerful in its own right – to build a map that simultaneously reveals the complexity of tumour architecture and multi-omic information about each and every cell.
Five years into their programme, with the pipeline now mature and consistently producing high-quality data, the team is beginning to delve into biological questions that their pipeline is uniquely positioned to answer. The pipeline’s development has relied on a global network of early-career researchers who, together with their multidisciplinary team leads, have cleared technological hurdles and advancing the technology to its current working state. We met with some of them to learn more.
The team’s mapping pipeline is illustrated here. After progressing through core analysis technologies, data flows and high-powered processing infrastructure, a sample is presented as a computerised 3D model, which is annotated with detailed, interactive information about each cell’s molecular characteristics.
All analysis begins with serial two-photon tomography (STPT) with a microscope that serially sections the sample into 15 µM slices, imaging four fluorescence channels simultaneously. During this step, physical sections are collected onto slides for further analyses throughout the rest of the pipeline. At the beginning of the team’s programme, this collection phase was a labour-intensive, manual process. “A major advancement of our workflow has been automating the collection of these sections,” explains Claire Mulvey, a principal scientific associate based at the CRUK Cambridge Institute, UK, who has been heavily involved in the STPT optimisation. “This automated section capture can now collect hundreds of slices from a single tissue sample and arrange them in the correct order, and is just one of the many ways we’ve upgraded the STPT instrument.”
STPT is an important step that produces a faithful 3D scaffold of the sample, upon which the results of all further analyses are annotated. This method has also allowed the team to explore many aspects of tumour architecture, including characterising the spatial relationships of cells within their environment, and it can be used to pinpoint physical regions of interest, such as areas showing vascular mimicry or metastasis, which can be further explored with higher-resolution imaging techniques.
To produce ultra-high-resolution proteomic maps of specific regions of interest, samples can be cut into even thinner 2 µM slices for imaging mass cytometry (IMC) – a combination of mass cytometry and immunohistochemistry with high-frequency laser ablation, which can simultaneously image as many as 40 proteins and their modifications. IMC was originally pioneered for 2D imaging, but in late 2021, the team published a landmark dataset showcasing how the team optimised the technique for 3D image analysis. “The spatial arrangement of cells in a tumour is immediately clear when you map them in 3D,” says Laura Kütt, PhD student at the University of Zurich, Switzerland, and lead author of the study. “With our 3D model, we could detect a cluster of epithelial cells that was potentially an early metastatic protrusion, which wouldn’t have been possible in 2D.”
IMC is an incredibly powerful technique, but challenges such as processing speed and throughput can limit its suitability for answering questions requiring large sample sizes and high throughput. To expand the pipeline’s applicability, the team has developed a new high-dimensional imaging approach: hyperplexed immunofluorescence imaging (HIFI). “HIFI is extremely gentle on tissue – meaning even very fragile samples can be investigated – and can be used to identify all major cell types in a tumour substructure, including their phenotype and localisation with respect to the wider tumour architecture,” says Spencer Watson, postdoctoral researcher at the University of Lausanne and the Ludwig Lausanne Branch, Switzerland.
HIFI, still a relatively novel technique, has yet to be fully integrated into the IMAXT pipeline. But in the future, Spencer hopes to perform HIFI on the full collection of slides from STPT, to first answer important questions about how cell biology changes across the entire tumour architecture, then zoom in on areas of interest by using IMC.
A novel aspect of this pipeline is the ability to link data from separately performed single-cell analyses and survey sequencing to build comprehensive expression profiles for each cell. Since the start of their programme, the team has optimised their pipeline of single-cell techniques, including optimising protocols, developing models to link RNA- and DNA-sequencing data, enhancing direct library preparation for single-cell whole-genome sequencing and applying single-cell profiling to clinical samples. “We’ve used these methods to examine tumour evolution and chromosomal structural aberrations at the single-cell level, and learn more about the underlying biology,” says Ciara O’Flanagan, a research associate based at BC Cancer, Vancouver, Canada. For example, matching cells’ single-cell DNA- and RNA-sequencing data has identified clonal populations with differential gene expression in triple-negative breast cancer, pinpointing new avenues of exploration to better understand the disease. “Integrating single-cell sequencing data with spatial transcriptomic profiling technologies reveals novel cell-cell interactions,” Ciara notes.
To map gene expression in space, two core technologies are used: expansion sequencing (ExSeq, a technique covered in last year’s report), which physically expands a sample to detect the subcellular localisation of specific RNA transcripts, and MERFISH (multiplexed error-robust fluorescence in situ hybridisation), which can achieve near-genome-wide, spatially resolved RNA profiling of individual cells with high accuracy and detection efficiency. “Profiling the transcriptomes of individual cells in their native environment allows one to uncover spatial gradients in gene expression across whole tissues, the spatial distribution of cell types, and even the subcellular localisation of DNA and transcripts,” says Leonardo Sepulveda Duran, a postdoctoral fellow based at Harvard University, in Boston, US.
Although each of the techniques in the pipeline is valuable in itself, “their integration gives an unprecedented view of the spatial dissemination of cancer cells in their surrounding environment,” according to New York-based research fellow Ignacio Vazquez-Garcia (Memorial Sloan Kettering Cancer Center and Columbia University). Ignacio’s role is to integrate disaggregated single-cell sequencing data with in situ measurements and images. “3D molecular mapping poses a number of challenges that we’ve managed to overcome so far. A computational challenge we’re currently addressing is how to bridge between modalities so that we can resolve spatial patterns across the tumour at the single-cell level.”
To build the final map, all spatial and single-cell data must be annotated on the 3D scaffold of the sample produced via STPT, but this process is challenging because each technique captures data with different characteristics and formats. Drawing inspiration from astronomical surveys, in which multi-wavelength images and data are stitched together to produce accurate maps of the sky, University of Cambridge’s Eduardo Gonzales Solares, a senior research associate with a background in physics and astronomy, has helped overcome this challenge. “On several occasions, we’ve asked ourselves: how did we solve this problem in astronomy?” says Eduardo. The team has adopted a technique that develops a ‘star field’ for each tissue sample, using agarose-bead reference points in the STPT analysis. “We wouldn’t be able to achieve such accurate image registration without this crucial step,” Eduardo elaborates.
Because a single sample analysed on the pipeline produces multiple terabytes of data, another major hurdle faced by the team is the phenomenal volume of data produced. “It’s not plausible to move such large quantities of data around the world,” Eduardo says. “Not everyone has access to high-end workstations, expensive software or local storage capacity.” Therefore, instead of bringing data to local computers with limited power, the team has developed automated workflows and analysis pipelines, as well as a new cloud-based compute lab allowing investigators to access data, run software and interpret results on demand. “We call it the IMAXT cloud,” Eduardo says. “It’s provided a new environment for scientific research and collaboration across the team.”
A final piece of the team’s puzzle is how to interact with the rich data produced by the pipeline. Although computers can handle such large amounts of information, initial exploration and interpretation of raw results by a human user is an indispensable practice for good science. To facilitate this process, the team has developed Project Theia, the world’s first virtual-reality cancer laboratory, in which users can ‘fly’ through and interact with their data in 3D, launch complex analyses and even extend analysis to 4D timeseries. Theia is planned to be released later this year and will equip researchers with an entirely new tool to investigate cancer biology.
“Now that our workflows are fully established, we can concentrate on using this pipeline of advanced techniques to answer key questions in tumour biology,” says Claire. In particular, the team is starting to use their pipeline to provide new ways to explore the poorly understood process of metastasis. Researchers have faced difficulties in studying the spread of cancer cells and in measuring metastasis-determining factors in situ long before the metastasis is detectable. The team’s integrated pipeline, with its ability to detect and investigate single cells within entire organs, should enable exploration of metastasis in unprecedented detail.
The team is also looking to understand the role of the immune system during tumour development, leveraging the pipeline’s ability to detect tumour populations and local microenvironments in 3D to study the dynamic spatial selection and pruning of cancer cells under the selective pressure of the immune system. “Our aim is to identify stereotypical models of spatial cancer growth in 3D and how they influence clone competition and immune containment of the tumour,” says Ignacio.
Looking ahead, the team continues to optimise what they can achieve with their pipeline. ‘Tiger teams’ – specialist multidisciplinary groups originally named by NASA – have been formed across the consortium to address exceptionally difficult technical challenges. Spencer leads a tiger team focused on spatial proteomics and how to derive biologically relevant findings from detailed information on the 3D spatial organisation of a tumour. Conversations within this group ultimately helped achieve the development of HIFI.
“A great deal of our studies have been enabled by the highly collaborative environment across our consortium,” Spencer says. “I don’t think anything we do at this level of research would be possible without team science like this. Being part of this Cancer Grand Challenge has improved my work and the scope of my goals, just by being around the ambitious nature of the whole project. This kind of ambition is infectious and inspires us all to push harder.”
The IMAXT team is funded by Cancer Research UK.
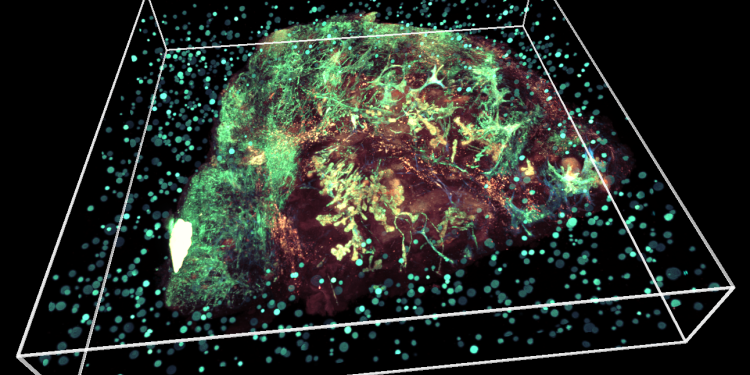
Written by Emily Farthing. Image: 3D tumour sample; credit: Eduardo Gonzales Solares and the IMAXT team. This article was originally included in our annual progress magazine Discover: A year of scientific creativity.
New findings from team eDyNAmiC
In this article, learn more about how IMAXT and Rosetta exploited new and existing technologies to propel spatial biology research.